One of the many shock waves created in COVID-19’s wake is a flood of companies employing artificial intelligence (AI) to quickly develop a safe and effective vaccine.
AI is a broad term that encompasses many technologies. The AI subfield that holds the most promise for vaccine development is machine learning (ML).
ML is an AI technique that can be used to design and train software algorithms to learn from and act on data. Software developers can use ML to create an algorithm that is ‘locked’ so that its function does not change, or ‘adaptive’ so its behavior can change over time based on new data.
The past several years have seen a revolution in ML. New hardware, especially Graphics Processing Units (GPUs), and the algorithms associated with “deep learning” have enabled computers to surpass humans in certain pattern recognition exercises for the first time.
For a basic overview on the AI/ML different terms, please see Karen Hao’s article “What is Machine Learning?” from the MIT Tech Review, available here.) Deep learning is a subfield of machine learning.

Source: https://www.edureka.co/blog/ai-vs-machine-learning-vs-deep-learning/
U.S. Patent No. 10,196,427
The patent’s author is Jacob E. Glanville. He is Co-Founder, CEO and President of DistributedBio. He and his teams apply computationally-guided immunoengineering methods to enable a new generation of monoclonal antibody discovery and universal vaccine design. The patent he wrote was assigned to his company, and it is this invention that appears to be the foundation for DistribedBio’s technology and success.
Dr. Glanville was featured in the Netflix documentary “Pandemic.” Following his path from the initial patent award in 2013 to his high-profile role in the quest for a vaccine against COVID-19 illustrates both the major advances in technology in recent years and the speed of development that such new technologies has brought about.
The patent is described as, “Epitope focusing by variable effective antigen surface concentration.” And the invention, “provides compositions and methods for the generation of an antibody or immunogenic composition, such as a vaccine, through epitope focusing by variable effective antigen surface concentration.”
The disclosure and the abstract is described as an invention that relies heavily on “in silico bioinformatics.” This means scientific experiments or research using computer modeling or computer simulation for the science of collecting and analyzing complex biological data. The disclosure describes neural networks to “generate a map of the protein surfaces of a particular antigen” or to “generate an in silico library of antigenic variants.” The abstract describes one step of the invention as generating ‘in silico’ “a library of potential antigens for use in the immunogenic composition.”
“Claims” are an important part of any patent. For example, Claim 1 says a, “method for eliciting an immune response in a human subject, the method comprising: delivering at least six antigens to the human subject, wherein each of the at least six antigens comprises: a target epitope that is common to each of the at least six antigens; and one or more non-conserved regions that are outside of the target epitope; wherein the at least six antigens are delivered such that each individual antigen of the at least six antigens is delivered in an amount that is insufficient to be immunogenic to the human subject on its own, while the at least six antigens are delivered in a combined amount that is sufficient to generate an immune response to the target epitope in the human subject.”
Claim 1 should be noted for its description of the patent’s purpose and outcome instead of the actual mathematical model that was developed to create the technology. This is because patents related to ML and AI should focus on the inventive means of achieving a result rather than the underlying mathematical concepts.
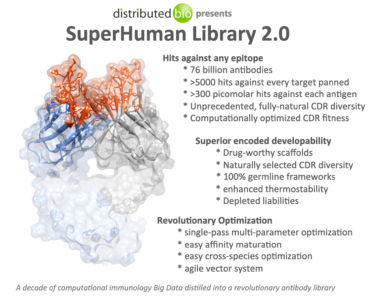
Claim 1 focuses on an “epitope.” An epitope, also known as antigenic determinant, is the part of an antigen that is recognized by the immune system, specifically by antibodies, B cells, or T cells. The image above describes the company’s Superhumn Library 2.0 as having 76 billion antibodies. Having so many antibodies to test against illustrates why ML is necessary to process so many iterations. The company calls this “antibody repertoire analysis.” New developments in chip design and the flexibility of cloud computing have made this type of work viable in recent years.
In 2018, DistributedBio announced an agreement with Pfizer Inc. to license its SuperHuman Platform 2.0 for the identification of novel antibodies for use as therapeutic agents.
Under the terms of the agreement, Distributed Bio agreed to transfer and license its platform to Pfizer, which it will use to perform screening activities against Pfizer-selected targets. Pfizer has also secured access to continued improvements to the SuperHuman Platform. Distributed Bio will receive an undisclosed annual licensing fee and future payments upon achievement of specified preclinical and clinical milestones.
“The SuperHuman Platform represents the culmination of a decade of our research in computational library design,” says Jacob Glanville, Co-Founder and Chief Science Officer of Distributed Bio. “From analyzing thousands of human immune systems with our machine learning AbGenesis platform, we have harvested from nature the rules of making exceptional therapeutic repertoires. The result is a library of 76 billion antibodies that contains over 5,000 hits against any antigen, including hundreds of picomolar binders, all thermostable, non-immunogenic, pre-screened by human blood and therapeutically developed in advance to avoid engineering delays downstream.